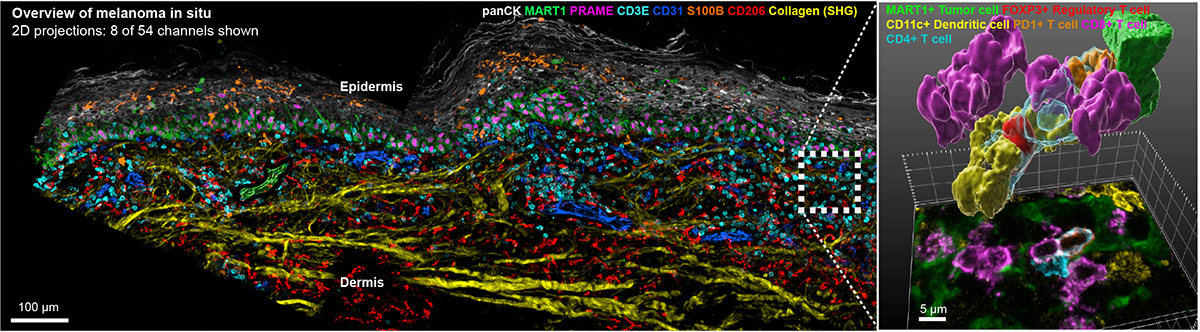
For 150 years, histopathological analysis of hematoxylin and eosin (H&E) stained specimens has been performed using tissue sections that are 4-5 μm thick. Contemporary “spatial biology” approaches have retained this format. The use of thin sections makes fine details of cells and organelles clearer (by preventing interference from out of focus light) but a report in the September issue of Nature Methods led by Ludwig Harvard’s Clarence Yapp, Ajit Nirmal, Sandro Santagata and Peter Sorger shows that nearly all cells and nuclei are incomplete in standard 5 μm sections. This compromises the accuracy of phenotyping and obscures functionally significant contacts between cells. By extending highly multiplexed spatial profiling to 3D imaging of 30-50 μm thick sections, the Harvard team has shown that it is possible to accurately determine the shapes of intact cells, analyze juxtracrine signaling events, and detect mitochondria, peroxisomes, secretory granules and other intracellular organelles. Analysis of cell–cell interactions in 3D shows that many immune cells within and at the margins of tumors are engaged in multiple (up to ten) simultaneous activating and inhibitory interactions with neighboring cells. 3D tissue imaging therefore promises to improve understanding of tumor, stromal and immune cell interaction in the presence and absence of immunotherapy. Precise mapping of T cell states and niches is a key goal for the next phase of this research.
Study data is freely available by clicking here.
Highly multiplexed 3D profiling of cell states and immune niches in human tumors
Nature Methods, 2025 September 29